Quansheng Du, PhD
Associate Professor
Dept. of Neuroscience & Regenerative Medicine
Mailing Address:
Dept. of Neuroscience & Regenerative Medicine
1120 15 St., Rm. CA3004
Medical College of Georgia at Augusta University
Augusta, GA 30912
E-mail: qdu@augusta.edu
Phone: 706-721-7370
Lab: Rm. CA3046
Education:
1985-1986 BS in Virology - Dept. of Virology and Molecular Biology, Wuhan University,
China
1992-1995 MS in Virology - Inst. of Virology, Wuhan University, China
1995-1998 PhD in Microbiology - Inst. of Virology, Wuhan University, China
Training:
1998-1999 Postdoctoral Training, Dept. of Pharmacology, University of Virginia
1999-2005 Postdoctoral Training, Center for Cell Signaling, University of Virginia
Research interests:
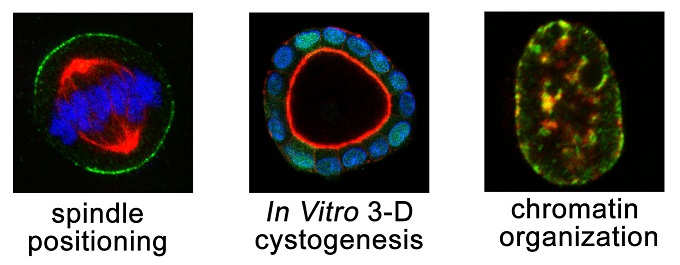
- Cell division. We are interested in the molecular mechanisms that regulate mitotic spindle organization and orientation in mammalian system. We are also interested in exploring the role of oriented cell division during epithelial morphogenesis and carcinogenesis.
- Cell polarity. We are interested in how polarity proteins are involved in cell-cell contact formation, cell polarization, cell-cell internalization and epithelial morphogenesis.
- Genome organization. We are interested in the molecular mechanisms underlying cell-cycle-regulated genome organization, focusing on the role of nuclear scaffold proteins.
- Stem cell. We are interested in molecular mechanisms underlying asymmetric cell division of stem cells. We are also interested in how polarity proteins are involved in stem cell fate determination.
Selected Publications:
- Du Q, Lehavi D, Faktor O, Qi Y, Chejanovsky N. “Isolation of an Apoptosis Suppressor Gene of the Spodoptera littoralis Nucleopolyhedrovirus.” Journal of Virology. (1999). 73:1278-85
- Du Q, Ren XR, Wang Q, Mei L, Xiong WC. “Inhibition of PYK2-induced cytoskeletal reorganization, PYK2 autophosphorylation, and focal adhesion targeting by FAK.” Journal of Cell Science, (2001). 114(16): 2977-87
- Du Q*, Stukenberg PT, Macara IG. “A mammalian Partner of inscuteable binds NuMA and regulates mitotic spindle organization.” Nature Cell Biology, (2001). 3: 1069-75
- Du Q*, Taylor L, Compton DA, Macara IG. “LGN blocks the ability of NuMA to bind and stabilize microtubules. A mechanism for mitotic spindle assembly regulation.” Current Biology, (2002). 12: 1928-33
- Wang Q#, Xie Y#, Du Q#, Wu X, Feng X, Mei L, McDonald JM, Xiong WC “Regulation of the formation of osteoclastic actin rings by PYK2 interacting with Gelsolin.” Journal of Cell Biology, (2003). 160: 565-75 (# these authors contribute equally to the work).
- Du Q, Macara IG. “Mammalian Pins is a conformational switch that links NuMA to heterotrimeric G proteins.” Cell, (2004). 119: 503-16
- Sans N, Wang PY, Du Q, Petralia RS, Wang YX, Nakka S, Blumer JB, Macara IG, Wenthold RJ. “mPins modulates PSD-95 and SAP102 trafficking and influences NMDA receptor surface expression.” Nature Cell Biology, (2005). 12: 1079-90
- Bowman SK, Neumüller RA, Novatchkova M, Du Q, Knoblich JA. “The Drosophila NuMA Homolog Mud Regulates Spindle Orientation in Asymmetric Cell Division.” Developmental Cell. (2006). 10(6): 731-42
- Zhu XJ, Wang CZ, Dai PG, Xie Y, Song N, Liu Y, Du Q, Mei L, Ding YQ, Xiong WC. “Myosin X regulates netrin receptors and functions in axonal path-finding.” Nature Cell Biology, (2007). 9(2):184-92
- Luo SW, Zhang C, Zhang B, Kim CH, Qiu YZ, Du Q, Mei L, Xiong WC. "Regulation of heterochromatin remodeling and myogenin expression during muscle differentiation by FAK interaction with MBD2". EMBO J. (2009). 28(17):2568-82
- Zheng Z, Zhu H, Wan Q, Liu J, Xiao Z, Siderovski DP, Du Q*. “LGN regulates spindle orientation during epithelial morphogenesis”, Journal of Cell Biology, (2010). 189: 275-88
- Hao Y, Du Q, Chen X, Zheng Z, Balsbaugh JL, Maitra S, Shabanowitz J, Hunt DF, Macara IG. “Par3 controls epithelial spindle orientation by aPKC-mediated phosphorylation of apical Pins”, Current Biology. (2010). 20(20):1809-18
- Zhu J, Wen W, Zheng Z, Shang Y, Wei Z, Xiao Z, Pan Z, Du Q*, Wang W*, Zhang M* “LGN/mInsc and LGN/NuMA Complex Structures Suggest Distinct Functions in Asymmetric Cell Division for the Par3/mInsc/LGN and Gai/LGN/NuMA Pathways”. Molecular Cell, (2011). 43:1-14
- Xiao Z, Wan Q, Du Q*, Zheng Z* “Galpha/LGN-mediated asymmetric spindle positioning does not lead to unequal cleavage of the mother cell in 3-D cultured MDCK cells”. Biochem Biophys Res Commun, (2012) 20;420(4):888-94
- Wan Q, Liu J, Zheng Z, Zhu H, Chu X, Dong Z, Huang S, Du Q* “Regulation of myosin activation during cell-cell contact formation by Par3-Lgl antagonism - Entosis without matrix detachment”. Molecular Biology of Cell, (2012). 23(11):2076-91, (Highlighted Article)
- Zheng Z, Wan Q, Liu J, Zhu H, Chu X, Du Q*. “Evidence for dynein and astral microtubule-mediated cortical release and transport of Gαi/LGN/NuMA complex in mitotic cells”. Molecular Biology of Cell, (2013). 24(7):901-13. (Highlighted Article)
- Slim CL, Lazaro-Dieguez F, Bijlard M, Toussaint MJ, de Bruin A, Du Q, Muesch A, van Ijzendoorn SC. “Par1b Induces Asymmetric Inheritance of Plasma Membrane Domains via LGN-Dependent Mitotic Spindle Orientation in Proliferating Hepatocytes”. PLoS Biology, (2013). 11(12): e1001739.
- Zheng Z#, Wan Q#, Meixiong G#, Du Q*. “Cell cycle-regulated membrane binding of NuMA contributes to efficient anaphase chromosome separation”. Molecular Biology of Cell, (2014). 25(5):606-19, PMID: 24371089 (# these authors contribute equally to the work)
- Zhu J, Shang Y, Wan Q, Xia Y, Chen J, Du Q, Zhang M. “Phosphorylation-dependent interaction between tumor suppressors Dlg and Lgl”. Cell Research, (2014). 24(4):451-63. PMID: 24513855
- Li Y, Wei Z, Yan Y, Wan Q, Du Q, Zhang M. “Structure of Crumbs tail in complex with the PALS1 PDZ-SH3-GK tandem reveals a highly specific assembly mechanism for the apical Crumbs complex”. Proc. Natl. Acad. Sci. U.S.A., (2014). 111(49):17444.
- Chu X, Chen X, Wan Q, Zheng Z, Du Q*. “NuMA Interacts with and Regulates Astrin at the Mitotic Spindle.” Journal of Biological Chemistry. 2016 Jul 26. pii: jbc.M116.724831
- Landin Malt A, Dailey Z, Holbrook-Rasmussen J, Zheng Y, Hogan A, Du Q, Lu X. (2019) Par3 is essential for the establishment of planar cell polarity of inner ear hair cells. Proc. Natl. Acad. Sci. U.S.A. doi: 10.1073/pnas.1816333116. Epub Feb 27.